Local AI Agent with RAG: Guide to Fact-Based Responses

Creating an AI assistant that is actually useful requires accurate information, and a plain LLM can't guarantee accuracy - knowledge is compressed into weights during training, so the model ends up guessing the details it doesn't really remember. We need something more.
That something is RAG - Retrieval-Augmented Generation. Instead of trusting the model to recall a fact, we retrieve the fact from our own data and hand it to the model as context. The model's job shrinks from "remember everything" to "summarize what I just gave you."
In this article we'll build exactly that: a local AI chat that solves a real-world problem based on actual data. If you're not familiar with concepts like vector search, embeddings, or how to build an AI agent, feel free to start with the previous articles.
Case study
A certain percentage of the population is sensitive to fermentable sugars (to put it simply). That means certain foods have to be avoided, and tracking which ones is tiresome. Current LLMs are decent at that but not always accurate - and "not always accurate" is a problem when the answer decides what someone eats. We can do better by grounding the assistant in actual science-based data, in this case the research paper: Presence of Fermentable Oligo-, Di-, Monosaccharides, and Polyols (FODMAPs) in commonly eaten foods.
The goal
The patient wants to know if drinking orange juice is safe - meaning the food doesn't contain an excess of any of the fermentable carbohydrates covered by FODMAP acronym. For this prototype our main goal is to return which substances are present, using the data we have in our dataset.
User
> Is orange juice low FODMAP?
Assistant
> Orange juice contains fructose (2.8g/100g).
Local model
Running an LLM locally sounds like a hassle and something that requires an extremely performant computer. But once the model is only responsible for summarizing retrieved data instead of remembering it, a small edge model (Gemma 4 E2B) is plenty. LM Studio lets you spin up an OpenAI-compatible HTTP server with one click, which makes it a perfect fit here. It also ships with nomic-embed-text-v1.5, which we'll use to turn text into vectors.
Turn data into embeddings
First we need the data in a saner format than PDF. I used Claude Cowork to convert it into CSV and dropped it into the repository so we don't have to worry about parsing. This is a simplified example of the CSV - a list of food items with the amount of each fermentable carbohydrate per 100 grams:
| Food Item | Fructose (g/100g) | Glucose (g/100g) | Fructan (g/100g) | Lactose (g/100g) | GOS (g/100g) | Polyol (g/100g) |
|---|---|---|---|---|---|---|
| Orange juice | 2.8 | 3.4 | 0 | 0 | 0 | 0 |
| Potato | 0.4 | 0.4 | 0 | 0 | 0 | 0 |
| Banana | 2.6 | 4.4 | 0.5 | 0 | 0 | 0 |
| Cola drinks | 3.4 | 3.5 | 0 | 0 | 0 | 0 |
| Tomato | 1.4 | 1.4 | 0.09 | 0 | 0 | 0 |
For every food in the CSV we build a short descriptive text blob, hash it, ask LM Studio for an embedding, and upsert into Postgres. Hashing the source text means we can re-run the seeder without re-embedding rows that haven't changed - embeddings aren't free, even locally.
# lib/fodmap/embedding_generator.rb (simplified)
def process(food)
text = build_source_text(food) # "Food: Banana\nPer 100g — fructose: 2.6g, ..."
hash = Digest::SHA256.hexdigest(text + @model_id)
existing = FodmapFood.find_by(name: food[:name])
return if existing && existing.source_hash == hash # idempotent re-run
vector = embed(text) # array of 768 floats from LM Studio
FodmapFood.upsert(
name: food[:name],
source_text: text,
source_hash: hash,
embedding: vector,
# ...numeric FODMAP columns
)
end
Table that speaks vectors
Postgres doesn't understand vectors by itself. We install the pgvector extension, which adds a vector column type and similarity operators. The migration pins the dimension to the embedding model (768 for nomic-embed-text-v1.5) - if you swap the model, you'll need a new column:
# db/migrate/..._create_fodmap_foods.rb
create_table :fodmap_foods do |t|
t.string :name, null: false
t.decimal :fructose_g, precision: 10, scale: 3
# ...more numeric columns
t.text :source_text, null: false
t.string :source_hash, null: false
t.column :embedding, "vector(768)"
end
add_index :fodmap_foods, :embedding, using: :ivfflat, opclass: :vector_cosine_ops
The neighbor gem turns similarity search into one line on the model:
class FodmapFood < ApplicationRecord
has_neighbors :embedding
scope :by_similarity, ->(vec) { nearest_neighbors(:embedding, vec, distance: "cosine") }
end
Embed the user's query the same way
To search, we embed the user's query into a vector and ask Postgres for the nearest neighbors. The critical rule: we have to use the same model as during ingestion. Different embedding models produce vectors in different coordinate systems, so mixing them silently breaks retrieval - the numbers still come back, they just don't mean anything.
# app/services/fodmap/query_embedder.rb (simplified)
def call
response = post(LmStudio::EMBEDDINGS_ENDPOINT, {
model: "text-embedding-nomic-embed-text-v1.5",
input: @text
})
JSON.parse(response.body).dig("data", 0, "embedding")
end
Don't trust blindly
Pure vector search is fuzzy by design. "Banana bread" and "banana" live close together, which is helpful for recall but dangerous for a factual lookup - we don't want to report banana's numbers when the user asked about banana bread. So we try cheap exact matches first and only fall back to embeddings:
# app/services/fodmap/food_matcher.rb (simplified)
def call
exact = FodmapFood.where("LOWER(name) = ?", @query.downcase)
return matched(exact.first, type: "exact") if exact.one?
vector = QueryEmbedder.call(@query)
candidates = FodmapFood.by_similarity(vector).limit(3).to_a
best = candidates.first
return not_found if best.neighbor_distance > 0.18 # too far → don't guess
return ambiguous(candidates) if close_second?(candidates) # two near-ties → ask
matched(best, type: "embedding")
end
Three safeguards here matter more than the retrieval itself:
- A distance ceiling - if nothing is close enough, return
not_foundinstead of the least-bad match. - An ambiguity gap - if the 2nd result is nearly as close as the 1st, flag it instead of silently picking one.
- Exact-match first - embeddings are a fallback, not a front door.
Extract ingredients first
There is one more small but important step before the lookup. The user usually doesn't type a clean database key. They ask real questions: "Can I eat pasta with garlic and tomato sauce?" or "Is banana bread OK?". Sending that whole sentence into a food matcher would make retrieval noisier than it needs to be.
So before matching, we ask the model to do a narrow extraction task: return only the food names explicitly mentioned by the user, as JSON. This keeps the lookup deterministic after the extraction step:
# app/services/fodmap/ingredient_extractor.rb (simplified)
SYSTEM_PROMPT = <<~PROMPT
Extract only the ingredient or food names explicitly mentioned.
Ignore quantities, measurements, cooking actions, and filler words.
Do not invent ingredients that are not present.
Reply with JSON only:
{"ingredients":["banana","milk","honey"]}
PROMPT
def call
content = lm_studio_complete(SYSTEM_PROMPT, @text)
JSON.parse(content).fetch("ingredients").map(&:downcase).uniq
rescue
[]
end
The tool then runs the matcher separately for each extracted ingredient and returns a structured payload:
# app/services/fodmap/ingredient_lookup_tool.rb (simplified)
def call
ingredients = IngredientExtractor.call(@text)
{
ingredients: ingredients.map do |ingredient|
result = FoodMatcher.call(ingredient)
serialize_match(result) # matched, ambiguous, or not_found
end
}
end
This makes the assistant more useful for normal questions without making the retrieval code responsible for understanding full sentences.
Give the LLM a tool
Now we need to connect our retrieval pipeline to the model. The cleanest way is tool calling: we describe our lookup as a function the model can invoke, and the model decides when to use it. Even edge models like Gemma 4 E2B are good enough at this - we write JSON in the OpenAI tool-calling schema, and the model handles the rest.
# app/services/chat/tool_definitions.rb
{
type: "function",
function: {
name: "ingredient_lookup",
description: "Look up FODMAP values for ingredients mentioned by the user.",
parameters: {
type: "object",
properties: { query: { type: "string" } },
required: ["query"]
}
}
}
Once the model calls our tool, we run the lookup and feed the result back as a tool message. The model then has the facts it needs to answer:
# app/services/chat/tool_call_runner.rb (simplified)
assistant_message["tool_calls"].each do |tool_call|
query = JSON.parse(tool_call["function"]["arguments"])["query"]
payload = Fodmap::IngredientLookupTool.call(query) # matched row(s) as JSON
tool_messages << {
"role" => "tool",
"tool_call_id" => tool_call["id"],
"name" => "ingredient_lookup",
"content" => payload.to_json
}
end
Keep the assistant in its lane
Because this is a food assistant, we also add a cheap classifier before the main chat flow. If the user asks about something unrelated, we don't need the full RAG pipeline at all:
# app/services/food_domain_classifier.rb (simplified)
SYSTEM_PROMPT = <<~PROMPT
Reply with exactly one word:
yes - if the message is about food, ingredients, diet, nutrition, or FODMAP
no - otherwise
PROMPT
def call
answer = lm_studio_complete(SYSTEM_PROMPT, @user_message)
answer.strip.downcase.start_with?("y")
rescue
true # fail open so a classifier error doesn't break the chat
end
This isn't a security boundary, but it is a useful product boundary. It keeps the assistant focused, avoids weird tool calls for unrelated questions, and makes the rest of the prompts simpler because the system has one job.
Facts are the facts
Once we give the model our facts, we have to make sure it actually sticks to them. Left to its own devices, an LLM will happily fill silence with plausible-sounding nonsense. The fix is a system prompt that sets clear rules and guardrails, so empty fields stay empty instead of getting invented:
LOOKUP_SUMMARY_SYSTEM_PROMPT = <<~PROMPT
You are summarizing FODMAP ingredient lookup results.
Use ONLY the raw tool data provided in the tool messages.
Mention when an ingredient is matched, ambiguous, or not found.
Treat missing values as unknown, not zero.
Do not invent serving thresholds or nutrition claims.
PROMPT
There are two model calls in the final path. The first one is the normal assistant call, where the model can decide whether to invoke ingredient_lookup. If it does call the tool, we don't just append the JSON to the current answer. We start a second, narrower summarization pass with a stricter system prompt and the raw tool messages:
# app/services/chat/assistant_reply.rb (simplified)
def call
return refusal unless FoodDomainClassifier.call(latest_user_message)
assistant_message = lm.complete(
response_messages,
tools: ToolDefinitions.all,
tool_choice: "auto"
)
tool_messages = ToolCallRunner.call(assistant_message)
return assistant_message["content"] if tool_messages.empty?
lm.stream([
{ role: "system", content: LOOKUP_SUMMARY_SYSTEM_PROMPT },
*conversation_messages,
assistant_message,
*tool_messages
])
end
That separation is important. The first call decides what information it needs. The second call is only allowed to explain the facts that came back from the tool.
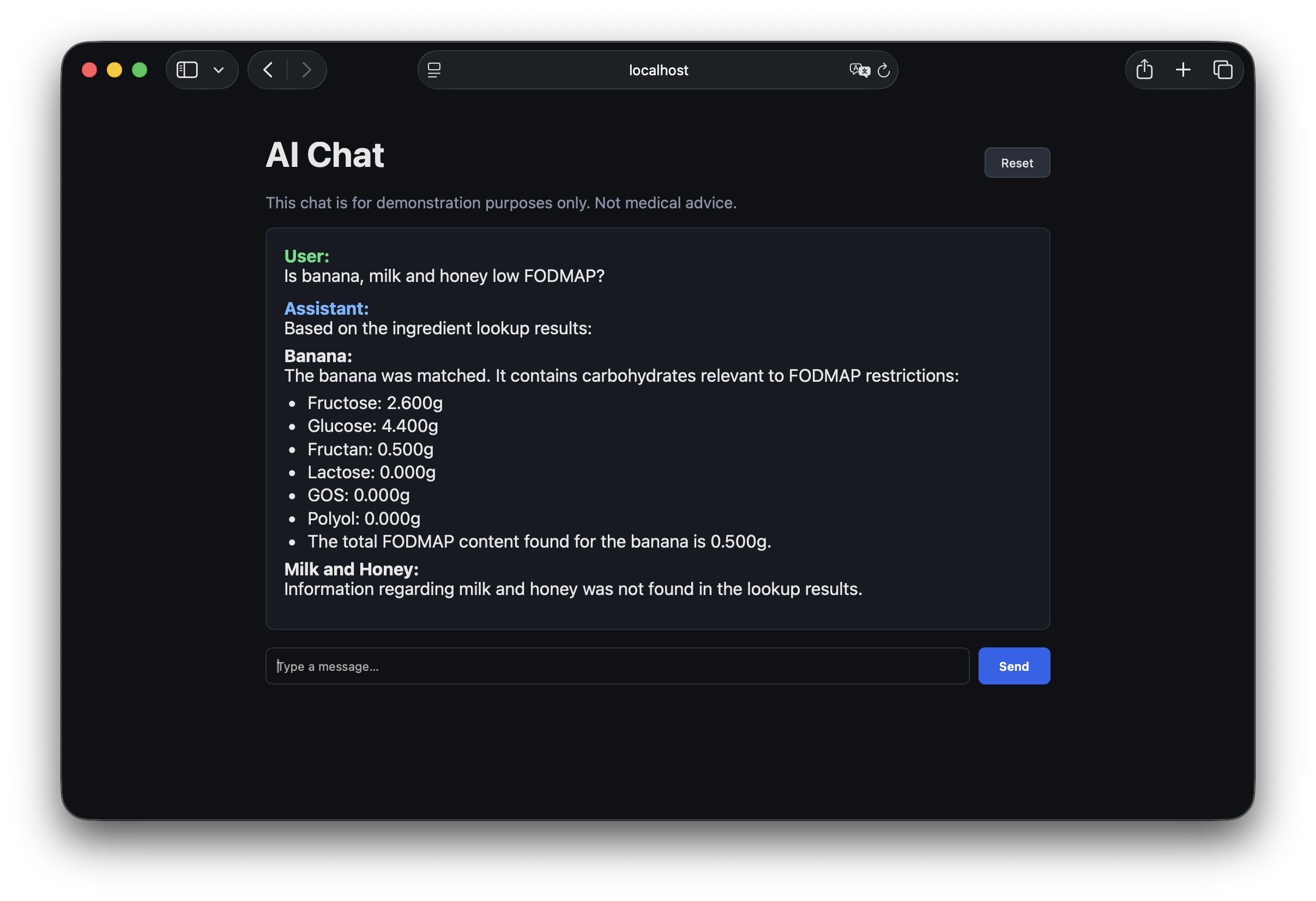
Wrapping up
The solution is far from perfect - but the model now keeps the rigorous scientific data intact instead of paraphrasing it into something plausible. From here we can expand in any direction: grow the dataset, tighten the guardrails, or teach the assistant to be more careful about edge cases. The most important thing is that we're grounded in facts.
The full source code is available on GitHub: visualitypl/fodmap-ai-agent.